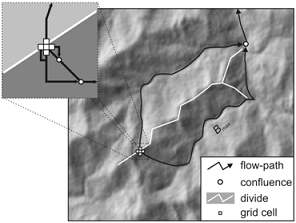
Maximum branch length (Bmax) is the longest branch length between a grid cell's flowpath and the flowpaths initiated at each of its neighbours. It can be conceptualized as the downslope distance that a volume of water that is split into two portions by a drainage divide would travel before reuniting.

If the two flowpaths of neighbouring grid cells do not intersect, Bmax is simply the flowpath length from the starting cell to its terminus at the edge of the grid or a cell with undefined flow direction (i.e. a pit cell either in a topographic depression or at the edge of a major body of water).
The pattern of Bmax derived from a DEM should be familiar to anyone who has interpreted upslope contributing area images. In fact, Bmax can be thought of as the complement of upslope contributing area. Whereas contributing area is greatest along valley bottoms and lowest at drainage divides, Bmax is greatest at divides and lowest along channels. The two topographic attributes are also distinguished by their units of measurements; Bmax is a length rather than an area. The presence of a major drainage divide between neighbouring grid cells is apparent in a Bmax image as a linear feature, often two grid cells wide, of relatively high values
Bmax is useful in the study of landscape structure, particularly with respect to drainage patterns. The index gives the relative significance of a specific location along a divide, with respect to the dispersion of materials across the landscape, in much the same way that stream ordering can be used to assess stream size.
The user must specify the name of a flow pointer grid derived using the D8 flow algorithm. In fact, the flow pointer raster could be derived using any non-dispersive flow algorithm that uses the same flow direction naming convention. The user must also specify whether or not to include diagonal neighbours in the calculation.
The following is an example of a Python script that uses this tool:
wd = pluginHost.getWorkingDirectory()
pointerFile = wd + "D8 pointer.dep"
outputFile = wd + "output.dep"
args = [pointerFile, outputFile]
pluginHost.runPlugin("BranchLength", args, False)
This is a Groovy script also using this tool:
def wd = pluginHost.getWorkingDirectory()
def pointerFile = wd + "D8 pointer.dep"
def outputFile = wd + "output.dep"
String[] args = [pointerFile, outputFile]
pluginHost.runPlugin("BranchLength", args, false)